A team of scientists has developed a computer simulation that would allow them to create an electronic toggle switch, expanding what a synthetic gene network designed for biocomputations can do.
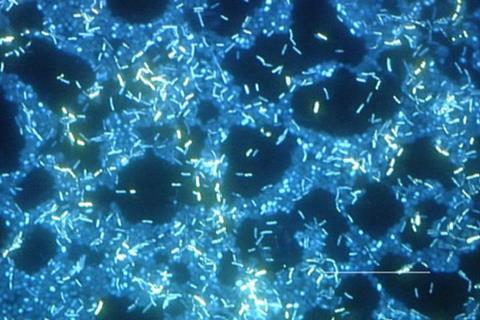
In a new paper, ‘‘An electrogenetic toggle switch model’, appearing in Microbial Biotechnology, an Applied Microbiology International publication, Lewis Grozinger and his team at Newcastle University used a computer simulation to find the principles for coordinating an electronic and bacterial system to create a toggle switch, potentially making it easier to control and measure a gene network in real time.
“Bacteria have already been engineered to host synthetic gene networks which can be designed to run biocomputations. The typical inputs and outputs of the gene networks are chemical, but we want to expand the types of input and output available for biocomputation to include electronic signals,” he told The Microbiologist.
“We expect that electronic signals linked to gene networks will make them easier to control and measure than chemical inputs and outputs, as well as offering a way to control and measure in ‘real time’ - something that traditionally synthetic biology has struggled with.
“Having gene networks linked to electrical signals also allows them to interface the bacterial hosts to external electronic circuits, which is something that fits well with the increasing trend of automation in the biological sciences.”
Designing the simulation
The team designed the simulation using an example of a very early synthetic gene network called the genetic toggle switch, hosted by a bacterium - this carries out a simple computation by remembering the last chemical input it received, and transforming it into a chemical output.
“We wanted to know if this kind of switch could be built to take an electronic input and produce an electronic output - it was not obvious that it should. So we modified the original design and simulated its operation,” Lewis said.
“We used computational simulations to find which parameters of the gene network would result in a functional switch. We called this new system the ‘electrogenetic toggle switch’.
“This means that it is in principle possible to build synthetic gene networks that are both controlled and monitored electronically in living bacteria.
“Here we used a very simple switch as an example, but these same computational simulations could be used to design any number of more complex gene networks. The bacteria and electronics then could coordinate the overall behaviour of the system by communicating in both directions, something that has not been considered in previous work.”
The next step, he said, is to use the simulation results and their expertise in synthetic biology to build and test the electrogenetic toggle switch in the bacteria Geobacter sulfurreducens, as well as other gene networks.
“If successful, this will significantly expand the range of tools available for engineering biology, opening up new kinds of workflows where electronics - for example, personal computers - are tightly integrated with the synthetic biological systems under study,” Lewis said.
The study was authored by Lewis Grozinger, Elizabeth Heidrich and Ángel Goñi-Moreno. Lewis Grozinger was supported by a PhD studentship from the EPSRC and was working in collaboration between the ICOS group at Newcastle University and the Biocomputation lab at the CBGP (UPM-INIA/CSIC), which is a lab led by Ángel Goñi-Moreno.
Elizabeth Heidrich is a research fellow at Newcastle University’s school of engineering.
Topics
- Ángel Goñi-Moreno
- Applied Microbiology International
- Bacteria
- biocomputation
- CBGP
- Community
- Elizabeth Heidrich
- genetic toggle switch
- Geobacter sulfurreducens
- Innovation News
- Lewis Grozinger
- Microbial Biotechnology
- Microbial Genetics
- Newcastle University
- Research News
- Synthetic Biology
- synthetic gene networks
- UK & Rest of Europe





No comments yet