Scientists at Johns Hopkins Medicine have developed a technique to find protein fragments that best stimulate the immune system to recognize and attack the HIV virus.
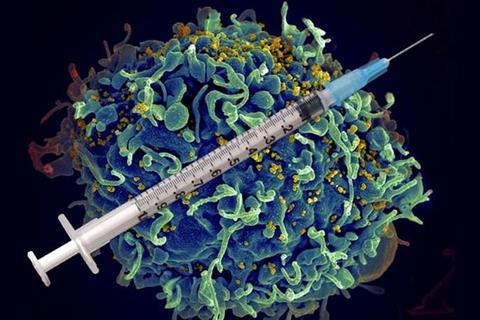
Currently, the WHO reports more than 38 million people globally live with the retrovirus, and each year, another 1 million new cases are diagnosed. While antiretroviral therapy helps keep HIV in check, patients must stay on their medication to prevent the development of AIDS.
Scientists have spent years trying to develop an effective HIV vaccine, but none have proven successful. Based on findings from a recently published study, a Johns Hopkins Medicine-led research team may have put science one step closer to that goal, as outlined in the Journal of Experimental Medicine.
Using a laboratory technique created at Johns Hopkins Medicine in 2010, the study researchers replicated the cellular environment in which specialized immune cells called antigen presenting cells (APCs) break down proteins derived from HIV and make them visible (“presented”) to the immune system’s frontline of defense, cells known as CD4+ T lymphocytes, or helper T cells.
Complex events
“Our simple method, called reductionist cell-free antigen processing, reproduces in a test tube the complex events that occur in the human immune system as a response to antigens, foreign invaders to the body such as viruses like HIV,” says senior study author Scheherazade Sadegh-Nasseri, Ph.D., professor of pathology at the Johns Hopkins University School of Medicine.
“When APCs chew up proteins from an antigen and present the fragments, known as antigenic epitopes, on their surface, the epitopes become visible to helper T cells and initiate an immune response.”
“If we can identify which epitopes are ‘immunodominant’ — the ones that elicit the strongest immune system response to the virus — then we may have the essential ingredients for the long-sought recipe to make an effective HIV vaccine,” explains Sadegh-Nasseri.
Lock and key
Epitopes that are immunodominant have structures that uniquely fit like a lock and key with cell-surface proteins on APCs known as major histocompatibility molecules, or MHCs.
“If you think of an HIV epitope as a hot dog and the MHC as a bun, the ‘meal’ is what gets presented to CD4+ T cells,” says lead study author Srona Sengupta, an M.D./Ph.D. candidate in immunology at the Johns Hopkins University School of Medicine. “T cells that can recognize the HIV epitope-MHC complex as foreign become activated and signal B cells — a different type of immune cell that produces antibodies, in this case, specific to HIV. Antibodies bind to the virus, destroying already infected cells or preventing HIV from entering uninfected ones — the key functions of an effective vaccine.”
Sadegh-Nasseri says previous efforts to map and identify the desired immunodominant epitopes have proven unreliable.
“Traditional methods use a ‘brute-force’ system where synthetic peptides representing portions of real HIV proteins are tested in the hopes that some will stimulate an immune response and direct researchers to the epitopes needed for vaccine development,” says Sadegh-Nasseri. “Not only is this strategy hit or miss, but the method doesn’t allow for the real-world chemical and molecular interactions that can impact how epitopes are produced and function.”
This, she explains, is a major reason why an effective HIV vaccine remains elusive.
Real-world interactions
“Our cell-free antigen processing system replicates how epitopes are actually processed in the APC’s cellular environment and become presented, including any influencing factors that may come into play,” says Sadegh-Nasseri.
“This enabled us to study nearly the entire HIV proteome [all of the proteins produced by the virus] and distinctly identify epitopes that are selected for presentation to CD4+ T cells by a chaperone protein called HLA-DM,” says Sengupta. “That’s important because we know that HIV epitopes processed and edited by HLA-DM are immunodominant.”
Sengupta adds that 35 epitopes identified in the recent studies were previously unknown.
The researchers say that their analysis using the cell-free antigen processing system revealed three important findings:
The epitopes identified are indeed generated in humans who are HIV positive and lead to the development of memory CD4+ T cells (the immune cells that remember an antigen for future encounters). The processing system can be very useful in predicting which parts of HIV protein antigens may yield the immunodominant epitopes that can be included in new vaccines.
The system’s use of full-length natural proteins ensures that the impacts of any cellular environmental influences (such as those causing modifications of viral epitopes after infected host cells have produced them) are taken into account.
Current analysis technologies lack such abilities, say Sadegh-Nasseri and Sengupta.
Surprising results
“Interestingly, we identified several epitopes that were modified by sugar groups, a potentially important finding for vaccine developers to know, but one that traditional analysis would have missed,” says Sengupta.
Sadegh-Nasseri and Sengupta say that their team will continue to refine the immunodominant epitope identification system and use the data from future analyses to enhance the ability of vaccine developers to design robust and effective protective measures against not only HIV, but also SARS-CoV-2 (the virus that causes COVID-19) and other viral pathogens.
Along with Sadegh-Nasseri and Sengupta, the members of the study team from Johns Hopkins Medicine and Johns Hopkins University are Nathan Board, Tatiana Boronina, Robert Cole, Madison Reed, Kevin Shenderov, co-senior author Robert Siliciano, Janet Siliciano, Andrew Timmons, Robin Welsh, Weiming Yang and Josephine Zhang. The team also includes Steven Deeks and Rebecca Hoh from the University of California San Francisco, and Aeryon Kim from Amgen Inc.
The study authors report no conflicts of interest.






No comments yet