Ashley Ward of SRUC and Sony Malhotra of STFC explain how an innovative collaboration explores how computational approaches could be used to detect non-O157 STEC that are likely to cause disease.
Shiga toxin-producing Escherichia coli (STEC) has long been known to pose a risk to human health via food sources. This group of bacteria are zoonotic, often transmitted by cattle, sheep, goats and deer, for which they cause no symptomatic disease.
Transmission to humans occurs via contact with the animals, through contaminated food or water, or from another infected person. Infection with just 10-100 organisms has serious consequences for human health and can cause serious gastroenteritis, or in severe cases, life-threatening haemolytic uremic syndrome.
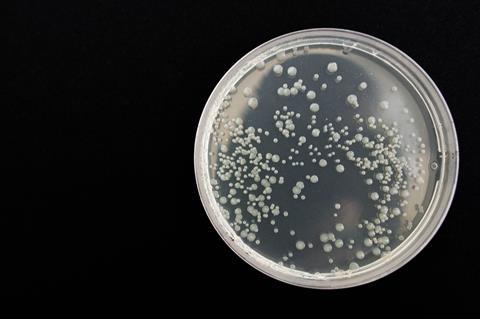
Monitoring for zoonotic threats at the site of primary food production enables detection of STEC, highlighting where there are risks of outbreaks. At the frontline of the surveillance system, diagnostic laboratories use a combination of culture- and molecular-based detection methods.
There are a variety of STEC serotypes and diagnostic labs screen for those that have been the most common cause of major outbreaks, using a risk-based framework. It is based on carriage of known virulence factors and works well for STEC serotypes like O157 and O26. However, many other STEC serotypes exist that are genetically very diverse, with only some able to cause severe human disease. The question is how to detect those that still have the potential to cause disease but that don’t fit into the current risk framework.
Recent outbreak
In the UK the incidence of non-O157 STEC disease cases is almost as high as O157. However, their prevalence is probably under-estimated because of difficulties with identification. This was highlighted in the UK during the winter 2023-2024 outbreak of STEC serotype O145 associated with cheese made from raw cows milk, for which the challenges in detection were noted by the producer. This outbreak caused 36 cases across the UK, including one fatality.
Determining STEC which are likely to cause disease is therefore a priority, to help us avoid serious ‘food safety incidents’. STEC have evolved relatively recently and continue to do so, with multiple genetic elements and mechanisms involved. The emergence of isolates with extensive genetic diversity generates new challenges beyond current diagnostic and surveillance capabilities, posing a risk to human health.
Addressing the challenge
Researchers at Scotland’s Rural College (SRUC), the Science and Technology Facilities Council (STFC) with an industry partner, Primer Design, have been exploring how computational approaches could be used to detect non-O157 STEC.
Since STEC genetic variation occurs both between and within genes, this group used two relevant approaches to see if the variation could be used as a predictor of disease, and in turn could be a good target for molecular detection strategies.
They collected a set of non-O157 STEC whole genome sequences representing clinical, food, and environmental isolates. For genetic variation within genes, they tested whether predicted protein function could be used as an indicator for pathogenic potential.
Protein pair
The well-characterised protein pair, Intimin (Eae) and the translocated intimin receptor (Tir) that facilitate host interactions by forming a very tight interaction, provided a good case study. The group modelled protein structures using the AI-powered tool, AlphaFold, and then protein function with STFC tools.
For variation between genes, a whole genome approach identified a group of genes more frequently carried by clinical isolates, so potentially a predictor of disease. This family are termed the Serine Protease Auto Transporters of Enterobacteriaceae (SPATE) and have been characterised as virulence factors for other types of disease caused by E. coli, such as urinary tract infections. Together, both sets of information were used by Primer Design to see if they could be incorporated into a molecular detection strategy.
For life-threatening pathogens like STEC, decision-making in clinical settings needs to be swift and efficient, while surveillance programmes need to be broad and inclusive to detect emerging threats. This project shows how an understanding of genomic variation can provide the fine level of detail that is needed to detect these evolving pathogens with the aim of reducing serious food safety incidents.
This project was funded through the Food Safety Research Network (FSRN - managed by the Quadram, a BBSRC network) and involves SRUC (Scotland’s Rural College), Science and Technology Facilities Council (STFC) and commercial partner Primer Design (part of Novacyt). To find out more, contact Professor Nicola Holden at nicola.holden@sruc.ac.uk or via LinkedIn.




No comments yet