Research has shed important new light on the enemies-turned-allies that allow bacteria to exchange genes, including those linked to antimicrobial resistance (AMR).
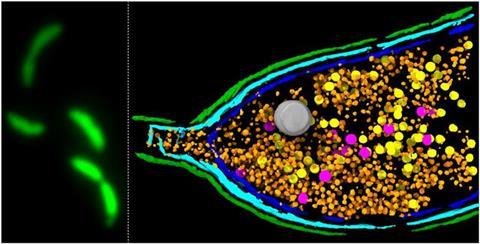
The insights, which expand our understanding of the major global health threat of AMR, came as John Innes Centre researchers investigated the curious phenomena of gene transfer agents (GTAs).
These gene-carrying particles look like bacteriophages (viruses that infect bacteria), but they have been domesticated from ancient viruses and put to beneficial use under the control of the bacterial host cell.
Acting as couriers, they take up parcels of host bacterial DNA and deliver them to neighbouring bacteria. This “selfless” sharing, known as horizontal gene transfer, can rapidly spread useful traits including genes that confer resistance to antibiotic drugs used to treat infections.
Host cell lysis
A crucial GTA life stage is host cell lysis: the breaking down of a host cell to release DNA-packed GTA particles. Previously, it was unclear how GTA particles escape their host bacterial cells.
In this study, which appears in Nature Microbiology, the team used a deep sequencing-based screening method to identify genes critical for GTA function in the model bacterium Caulobacter crescentus.
This identified a three-gene control hub, LypABC, encoding bacterial proteins. When these lypABC genes were deleted, bacteria could no longer lyse to release GTA particles. In contrast, by overexpressing the lypABC hub they obtained a very high proportion of lysing cells. Together, these experiments identified LypABC as a control mechanism for GTA-mediated cell lysis.
Repurposed system
Surprisingly, LypABC resembles a bacterial anti-phage immune system since it contains protein domains which are typically required for defence against viruses. However, this collaborative effort between the John Innes Centre, the University of York, and the Rowland Institute at Harvard, suggests it has been repurposed to release GTA particles for gene transfer.
They also identified a regulatory protein which is required for strict control of both GTA activation and GTA-mediated lysis. This control is important as misregulation of LypABC is highly toxic to bacterial cells.
In highlighting the plasticity of bacterial domains, the study advances fundamental knowledge of how gene transfer occurs between bacterial cells and offers an important clue to understanding how AMR occurs.
Sharing DNA
First author of the study Dr Emma Banks, a Royal Commission for the Exhibition of 1851 Research Fellow, said: “What’s particularly interesting is that LypABC looks like an immune system, yet bacteria are using it to release GTA particles. It suggests that immune systems can be repurposed to help bacteria share DNA with each other - a process that can contribute to the spread of antibiotic resistance.”
The next step for the research is to discover how the LypABC control hub is activated and how it functions to control the rupture of bacterial cells and release of GTA particles.
“A bacterial CARD-NLR-like immune system controls the release of gene transfer agents”, appears in Nature Microbiology.




No comments yet