Engineered microorganisms are widely used in industrial biotechnology and biopharmaceutical applications, including the production of biofuels, sustainable chemicals, and therapeutic compounds. However, concerns remain regarding the unintended environmental release and uncontrolled proliferation of genetically engineered microbes. For this reason, biocontainment technologies, which are designed to prevent microorganisms from surviving outside controlled environments, have become increasingly important in both academia and industry.
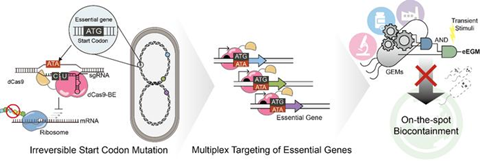
Conventional biocontainment strategies have relied on auxotrophy-based approaches, toxin–antitoxin systems, or DNA cleavage-based technologies such as CRISPR-Cas9. However, these methods often suffer from environmental dependency, genetic instability, and the risk of unintended mutations and cellular stress caused by DNA double-strand breaks.
In particular, DNA cleavage-based systems may compromise genomic stability and allow certain mutant cells to escape survival control. In addition, CRISPR interference (CRISPRi)-based systems are inherently reversible, posing challenges for achieving complete and permanent control of cell viability.
Precise nucleotide changes
In this study, published in Nucleic Acids Research, the research team employed a CRISPR-dCas9-based base editing system capable of introducing precise nucleotide changes without inducing DNA double-strand breaks. The researchers specifically targeted the start codons of essential genes and irreversibly disrupted their function, thereby permanently blocking cell survival. In simple terms, the system functions by permanently switching off the “power buttons” required for microbial survival.
Because this strategy does not directly cut DNA strands, it significantly reduces cellular damage and the risk of unwanted mutations compared with conventional CRISPR systems. Furthermore, the team implemented a multiplexed targeting strategy that simultaneously edits multiple essential genes. This approach dramatically reduced escape frequency, in which rare mutant cells survive despite containment measures.
The researchers also demonstrated that even short-term induction of the system was sufficient to irreversibly eliminate cell viability, highlighting the efficiency and robustness of the technology compared with conventional systems that require continuous expression. The team expects that this platform could serve as a next-generation biocontainment strategy for safer and more reliable control of engineered microorganisms.
Irreversible biocontainment system
The research team expects that this irreversible biocontainment system will provide an important technological foundation for improving the safety of genetically engineered microorganisms that may be released into the environment. The technology also presents broad applicability in industrial biotechnology and biopharmaceutical fields.
For example, the system could be applied as a biological safety mechanism in microbial production processes used for biofuels, biodegradable plastics, and high-value chemicals. In addition, it may serve as a controllable safety switch for live biotherapeutics and next-generation cell-based medicines operating inside the human body. The researchers anticipate that this technology will contribute not only to safer biomanufacturing platforms for industry but also to improving public trust in the practical use of engineered microorganisms.
Next generation biosafety
Professor Sang Woo Seo stated, “This study presents a novel strategy for precise and irreversible control of microbial cell survival using base editing. We believe this technology has strong potential as a next-generation biosafety platform.”
First author Dr. Sung Won Cho is continuing research on synthetic biology-based microbial control technologies and next-generation genome engineering systems, with a focus on biosafety and biomanufacturing platforms. Co-first author TaeHyun Kim is conducting research in synthetic biology and microbial systems engineering and plans to further pursue advanced studies in next-generation biotechnology platforms.
This work was supported by the Bio & Medical Technology Development Program through the National Research Foundation of Korea (NRF), funded by the Ministry of Science and ICT (MSIT), and by an NRF grant funded by the Korean government (MSIT). This work was also supported by the Samsung Research Funding & Incubation Center of Samsung Electronics.
Topics
- Asia & Oceania
- Bacteria
- biocontainment
- Biosafety
- Clinical & Diagnostics
- CRISPR-dCas9
- DNA double-strand breaks
- Economic Equality
- Industrial Microbiology
- Medical Microbiology
- Microbial Biotechnology
- One Health
- Quality Control
- Research News
- Sang Woo Seo
- Seoul National University College of Engineering
- start codons
- Sung Won Cho






No comments yet