Multicellularity, the aggregation of unicellular organisms into multicellular structures, is a key biological phenomenon for understanding the evolution of life. Social amoebas, known as cellular slime molds, live as individual cells under normal conditions, but cooperate to form multicellular bodies when nutrients become scarce, making them excellent model organisms for studying multicellular development.
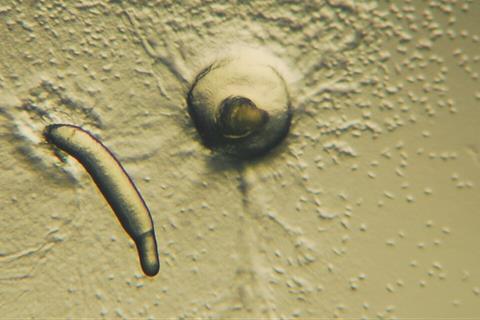
However, molecular genetic analyses have been largely limited to a single model species, Dictyostelium discoideum, making comparative studies across species difficult.
A research group led by Associate Professor Tetsuya Muramoto from the Faculty of Science, Toho University, has established a CRISPR genome editing technique that enables comparative analysis of the evolution of multicellularity across different species of social amoebas (cellular slime molds). Until now, genetic studies had been largely restricted to a single model species, limiting cross-species comparisons. These findings were published in the scientific journal Scientific Reports on February 5, 2026.
The research group expanded genome editing methods for social amoebas. Using a CRISPR vector applicable to multiple species, they successfully achieved gene modifications in several Dictyostelia species, ranging from ancestral to more morphologically complex groups. Furthermore, the co-introduction of donor DNA greatly improved editing efficiency, enabling gene disruption in species that had previously been challenging to genetically manipulate.
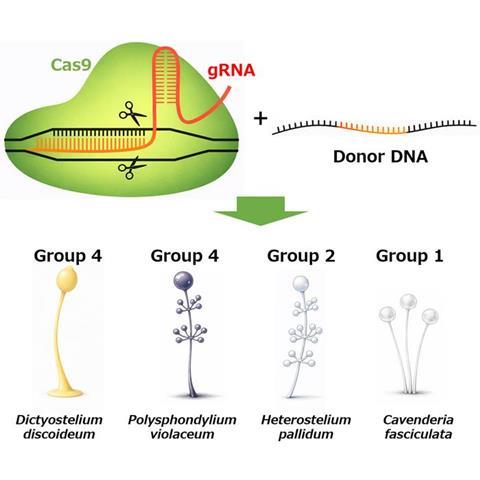
This study establishes a research platform that enables cross-species comparisons of how unicellular organisms cooperate to form multicellular structures. This is expected to significantly advance our understanding of the evolution of multicellularity and coordinated cell behavior.






No comments yet