A research group led by Project Professor Kazuyoshi Murata at the Exploratory Research Center on Life and Living Systems (ExCELLS) / National Institute for Physiological Sciences (NIPS), in collaboration with Senior Researcher Kenta Okamoto at Uppsala University and Professor Chantal Abergel at Aix-Marseille University, has successfully determined, for the first time in the world, the capsid (outer shell) structure of Melbournevirus—a member of the giant virus family—at a resolution of 4.4 Å using cryo-electron microscopy (cryo-EM).
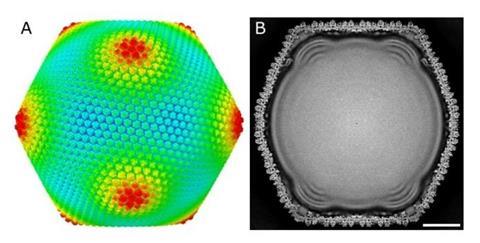
In this study, the researchers applied a “block-based reconstruction method” to the analysis of electron microscopy images, achieving a dramatic improvement in the resolution of the three-dimensional reconstruction (Figure 1).
This enabled them to elucidate the detailed arrangement of proteins constituting the massive capsid (250 nm in diameter) (Figure 2).
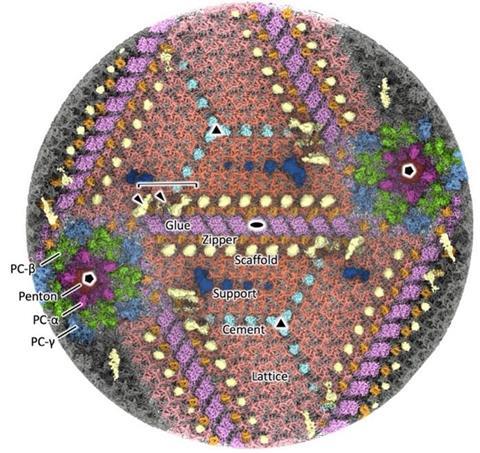
These findings contribute to understanding the fundamental principles by which large, uniform, and robust structures are formed from a limited set of proteins. In addition, the results are expected to advance our knowledge of viral evolution and infection mechanisms, and may also be applied to the design of inclusion compounds and drug delivery carriers.






No comments yet