Researchers at Liverpool School of Tropical Medicine (LSTM) have identified a bacterium strongly associated with noma disease, marking a major step towards earlier diagnosis and more effective treatment.
In a new study published in PLOS Neglected Tropical Diseases, a research team from LSTM, working with partners at the University of Liverpool, Médecins Sans Frontières and the Noma Children’s Hospital, Sokoto, Nigeria, used metagenomic sequencing and machine learning algorithms to analyse saliva samples from children with acute noma. They uncovered a previously undescribed species of Treponema bacteria, which may be responsible for the disease.
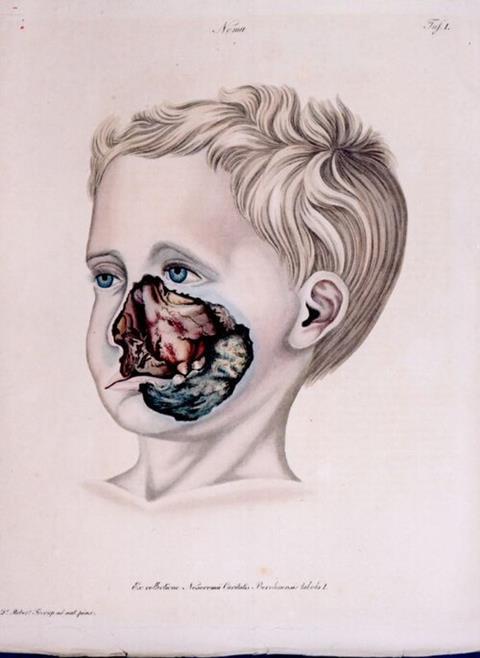
Noma is a rapidly progressing infection that begins as gingivitis and inflammation of the gums but goes on to destroy the tissues of the mouth and face. It primarily affects young children living in extreme poverty. Without treatment, it is fatal in up to 90% of cases, while survivors are left with severe disfigurement and lifelong complications.
Despite being formally classified as a neglected tropical disease by the World Health Organization in 2023, the underlying cause of noma has remained poorly understood.
Full bacterial community
Senior author on the study Professor Adam Roberts, from LSTM’s Centre for Neglected Tropical Diseases and a member of Applied Microbiology International’s One Health Advisory Group, said: “Noma is a disease that simply should not exist, but up until now we have lacked a detailed understanding of the microbes involved. By using metagenomic sequencing, we can now see the full bacterial community in unprecedented detail. This has allowed us to identify a species of Treponema bacteria strongly linked to the disease.”
READ MORE: Freshwater snails are an overlooked reservoir of AMR, study reveals
READ MORE: Tropical diseases: Global health meeting showcases resilience of field under fire
Importantly, for diagnosis and treatment, Treponema bacteria were detected in the early stages of the disease, suggesting they may play a key role in triggering progression from gingivitis to noma.
Further analysis showed that the bacteria do not carry known antibiotic resistance genes, suggesting that they could be effectively targeted with existing antibiotic treatments if detected early enough. Currently, most cases are identified too late, when severe tissue damage has already occurred.
Oral microbiome
The saliva samples also showed that noma had disrupted the oral microbiome, changing the balance of bacteria in the mouth.
Joint lead author Dr Richard Goodman said: “We found that the composition of bacteria in children with noma is completely disrupted. Some bacteria, such as those in the Treponema genus, appear to be more common, while others that are usually part of a healthy mouth almost disappear. That imbalance gives us important clues about how the disease starts and progresses.”
The findings open the door to the development of diagnostic tools that could detect noma at its earliest stages, before severe tissue damage occurs and when treatment is most effective.
Clearer picture
Joint lead author and PhD candidate Angus O’Ferrall said: “We now have a much clearer picture of the microbes linked to noma. This creates real opportunities for earlier detection, timely treatment and prevention by addressing the underlying microbial imbalance that drives the disease.”
By identifying, for the first time, a consistent microbial signature of noma, the study marks a major step forward in understanding a disease that has remained neglected for far too long, and brings renewed hope for effective prevention, diagnosis and treatment.




No comments yet