Anyone who has ever used a microscope knows that it takes time to bring a sample into sharp focus. Each time you move the slide, the image blurs, and you have to stop and carefully turn a knob to bring everything back into clear view. For scientists and clinicians, even if the motion is semi-automated, that time quickly adds up as they work with dozens or hundreds of samples.
Now a team of scientists at Caltech has developed an inexpensive robust fix for this problem that involves little more than a couple of LED lights and some physics-based processing. They describe the new autofocus technique, which they call Digital Defocus Aberration Interference (DAbI), in a paper published online on April 24 in the journal Nature Communications.
The lead authors of the paper are graduate students Haowen Zhou (MS ’24, PhD ’26) and Shi “Josh” Zhao (MS ’25), who completed the work in the lab of Changhuei Yang, the Thomas G. Myers Professor of Electrical Engineering, Bioengineering, and Medical Engineering at Caltech and a Heritage Medical Research Institute Investigator.
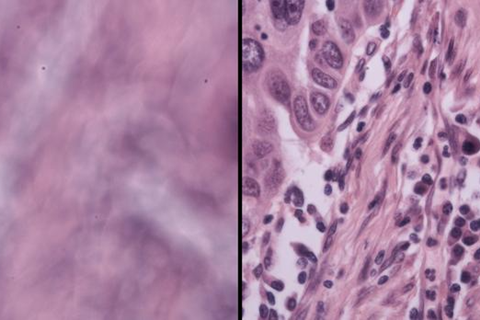
The underlying concept is fairly simple. When two LEDs illuminate a sample from slightly different angles, the combined signal derived from two photographs (one taken at each source of illumination) reveals a hidden fringe, or pattern of stripes. Those stripes change in a predictable way depending on how far the sample is from the focal point, the sweet spot where an image comes into focus. Therefore, a computer reading the stripes can tell the microscope how to correct for any blurriness in the image.
Accidental discovery
The discovery was somewhat accidental. “We were debugging for another project,” Zhou says. “When we summed up the contributions from the photos taken from two different locations, we found this fringe pattern. If we defocused more, the fringe would be much denser. If we defocused less, it would be more spread out. So, it seemed there was a strong correlation between the defocus value and the fringe density.”
The scientists have tested DAbI on six different types of microscopes—from basic compound light microscopes to more complex systems used for imaging living cells and tissues, or even thick 3D specimens—all with excellent results. When dealing with thin flat samples, DAbI kept images in focus across a range more than 400 times larger than the natural depth of focus of a basic microscope lens.
“Our technique, which is enabled by a physics-based observation, is reliable, high performance, and also very simple,” Zhou says. “This makes it useful and powerful for automated, high-throughput microscopy.”
Unique technique
Zhao adds that the DAbI technique is unique in that it can be used to locate the plane in which the focal point exists, even in thick 3D samples. Indeed, for thicker 3D samples up to 150 micrometers deep, DAbI achieved a range nearly 300 times larger than the natural limit in tests. “This offers truly robust autofocusing of 3D samples, which has never been possible with other techniques,” Zhao says.
The team has collaborated with other labs both on campus and elsewhere to test DAbI on a variety of samples, including ecological samples such as algae and bacteria, as well as embryological, oncological, and neurobiological samples.
“This work exemplifies how Caltech’s unique research ecosystem enables swift and close collaborations among the research groups here,” says Yang, who is also the executive officer for electrical engineering at Caltech.
Brain slices
To test the technique on brain slices and organoids, simple versions of organs that are grown in vitro, Zhou and Zhao reached out to the lab of Viviana Gradinaru (BS ’04), the Lois and Victor Troendle Professor of Neuroscience and Biological Engineering, director and Allen V. C. Davis and Lenabelle Davis Leadership Chair of the Richard N. Merkin Institute for Translational Research at Caltech, and a Howard Hughes Medical Research Institute Investigator. Gradinaru and postdoctoral scholar Yujie Fan helped the team prepare samples for testing, including brain slices ranging from 50 to 100 nanometers in thickness.
“Because of the complex 3D structures and irregular shapes of these thick samples, it’s really hard for the current microscope system to find the perfect focus automatically,” Fan explains. “DAbI performed exceptionally well even with thick biological samples.”
Fan adds that if DAbI can be integrated into standard microscope systems, it will be “a game changer,” turning a slow manual process into truly automated imaging, “not only saving researchers like me a massive amount of time, but also making imaging-based high-throughput screening of 3D tissue possible,” she says.
The paper is titled “Digital defocus aberration interference for automated optical microscopy.” Additional authors are Caltech graduate student Zhenyu Dong (MS ’25) and Oumeng Zhang, a former postdoctoral scholar from Yang’s group who is now at Texas A&M University. The work was supported by funding from the Rothenberg Innovation Initiative, the Heritage Medical Research Institute, a Caltech Chen Postdoc Innovator Grant, and a Caltech Schmidt Graduate Research Fellowship.






No comments yet