Here’s the reality: a stool report that reads “Blastocystis detected” still provokes strong reactions. Some clinicians worry and reach for antibiotics. Some laboratories add a note about “uncertain significance.” Patients search online and find polarised claims ranging from harmless commensal to stealth pathogen. The truth is more nuanced and more informative than those extremes. Blastocystis is one of the most frequently detected gut protists in humans and many animals. High prevalence does not mean pathogenicity, but it also does not exclude context-dependent effects. The question isn’t “good or bad?” but rather “which subtype, in which host, with what microbiome, and over what timeframe?”
Researchers should be cautious when designing and interpreting studies of Blastocystis. Subtype-blind assays, small cross-sectional samples, and inadequate control for antibiotics, travel, diet, and co-infections can inflate fragile associations. Several recent papers have prompted formal concerns from editors about mis-citation and over-interpretation; until standards mature, treat single-method signals as hypothesis-generating and publish protocols, raw data, and metadata alongside the results.
What exactly are we dealing with?
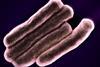
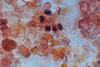
Blastocystis is a genus of anaerobic microbial eukaryotes that colonise the large intestine (Figure 1). It’s genetically diverse. Multiple subtypes (STs) circulate in humans and animals, and those STs likely differ in host range, ecological strategy, and interactions with resident microbes. That diversity matters for biology and for public health interpretation. We see Blastocystis often because it’s widespread in mammalian and avian hosts, transmits faecally–orally, and survives well enough in the environment for water and food to play a role. Add the wide variation in assay sensitivity, and detection in asymptomatic people is no surprise.

Prevalence is not destiny
Frequent detection in healthy groups challenges the idea of a straightforward pathogen. However, the claim that it is “always harmless” does not fit either. Associations with abdominal pain or IBS-like symptoms appear in some studies and disappear in others. When signals do emerge, they often fade after accounting for antibiotics, travel, co-infections, diet, or the broader microbiome. This indicates that context plays the most significant role.
Two details are important and often overlooked: quantity and duration. A binary positive does not indicate whether the load is negligible or dominant, or if colonisation is transient, intermittent, or persistent. Symptoms, if they appear, depend on a threshold of abundance maintained over time, not a single PCR result. Include subtype or strain differences, and you can understand why results conflict: two “positives” may be biologically very different.
The host backdrop also influences the outcome. Mucosal barrier integrity, bile acid metabolism, recent antimicrobial use, and diet can sway the gut ecosystem towards tolerance or irritation. In one person, Blastocystis might be a quiet tenant in a diverse community; in another, with antibiotics on board and a thinned mucus layer, it could intensify existing discomfort. Study design amplifies variability: clinic-based cohorts tend to include individuals already unwell, while cross-sectional snapshots cannot distinguish cause from consequence. Methodology adds further complexity; microscopy often underestimates; different primer sets have varying subtype blind spots; and shotgun metagenomics faces challenges at low abundance or high host DNA content.
What this truly means is that we should abandon the binary label. Replace it with questions that align with biology: which subtype or strain, at what load, for how long, in which host and microbiome context, and alongside what exposures? That framing is more honest for patients and more valuable for research, and it’s where longitudinal, subtype-aware, and quantitative studies will genuinely make a difference.

Subtypes and strain-level variation
Subtype-aware thinking is the missing component of the field. ST1–ST9 and beyond have been identified in humans, with ST1–ST4 being common in many regions. Some subtypes are widely shared across species, while others appear more host-specific. Within a subtype, strains can vary in gene content and surface features that influence colonisation, immune recognition, nutrient utilisation, and possibly interactions with bacteria. Two patients “with Blastocystis” may carry organisms with very different capabilities. Diagnostics that only determine presence or absence overlook significant details; incorporating subtype – and eventually strain – yields much clearer information for surveillance and research.
The microbiome context
Blastocystis lives inside a dense web of metabolic hand-offs. Bacteria, archaea, viruses, and other eukaryotes trade carbon and electrons; short-chain fatty acids, hydrogen, lactate, and bile acids move between partners. Where Blastocystis slots into this web likely depends on oxygen and nutrient gradients near the epithelium, access to mucins and dietary glycans, and which keystone bacteria are present. That’s why two people with a “positive” result can experience very different things.
Several cohorts report higher bacterial diversity in colonised individuals. It’s only an association, but it challenges the idea that Blastocystis is a universal marker of dysbiosis. Signals from culture and microscopy suggest that some strains feed on bacteria or their products. In contrast, others might benefit from cross-feeding with butyrate producers or methanogens via the hydrogen economy. Secreted enzymes – proteases and glycosidases – could alter local niches by thinning mucus or exposing new glycans, effects that might be neutral in a resilient community but irritating in a fragile one.
The host environment is important. If mucus barrier integrity is borderline, if bile acid signalling is already disrupted (after antibiotics or rapid weight loss), or if the diet is low in fermentable fibre, small changes in metabolite levels can feel significant. In a healthy, diverse microbiome, the presence of Blastocystis may go unnoticed. Medications that alter the microbiome (e.g., antibiotics, PPIs, metformin) can further shift the balance, not by “activating” Blastocystis directly, but by altering its environment.
These are all testable. Pair subtype-resolved, quantitative detection with longitudinal stool and mucosal sampling; measure SCFAs, bile acids, and inflammation markers; use co-culture and mini-bioreactors with keystone bacteria; run stable-isotope tracing to map who eats what; perturb with antibiotics, fibres, and bile acids. The goal isn’t to prove good or bad: it’s to find the conditions under which colonisation is clinically meaningful.
Diagnostics: what labs can do better today
Interpretation depends on methods. Light microscopy may miss positives. Broad-range PCR detects more but might under-detect specific subtypes. Shotgun metagenomics can identify Blastocystis reads, but sensitivity decreases at low abundance or when host DNA dominates. Reports should specify the method used, state detection limits, and indicate whether subtyping was performed. Instead of generic caveats, offer clear, neutral guidance: Blastocystis is often found in both healthy individuals and those with gastrointestinal symptoms; detection alone does not establish causation. Encourage clinicians to consider recent antibiotic use, travel history, co-pathogens, and symptom duration before deciding on treatment. When symptoms persist without an alternative explanation, recommend clinical correlation, possible retesting, or broader investigations rather than reflex therapy. Because interpretation rises or falls with methods, we need to be exacting. Recent manuscripts that drew formal concerns from editors show how methodological shortcuts and mis-citation can distort conclusions; single-method positives should not be presented as causal findings without subtyping, quantification, and covariate control.
Treat or watchful waiting?
Practice varies. Some clinicians treat symptomatic positives empirically; others adopt watchful waiting, especially when co-pathogens are absent and symptoms are mild. Randomised trials remain limited and small. Antibiotics can reduce or eliminate detection, but recurrence is common, and off-target microbiome effects complicate interpretation. If treating, document symptoms, co-infections, antibiotic history, and follow-up testing. If not treating, explain the rationale and set expectations for monitoring and re-evaluation.
One Health makes the difference: connecting people, animals, and environments
The reason a One Health perspective matters for Blastocystis is simple: the organism does not respect our disciplinary boundaries. Humans, livestock, companion animals, wildlife, water, soil, and food chains are part of one interconnected system. When you map detections across this system, patterns emerge that never become clear in studies that focus solely on humans. Subtypes that circulate among ruminants or poultry appear in nearby communities; seasonal increases due to irrigation or rainfall show up as changes in environmental load; household clustering reflects pet ownership and sanitation practices. What this truly means is that “Is Blastocystis present?” is the wrong question. The correct questions are where, in whom, in what subtype, and under which environmental conditions.
A practical One Health approach starts with mapping transmission routes across the farm-to-fork and waste-to-water pathways. On the food side, key points include primary production, manure management, irrigation water, processing facilities, retail produce, and household kitchens. On the waste side, consider septic tanks and sewers, wastewater treatment plants, rivers and coastal areas receiving effluent, and the re-use of biosolids. Also include animal interfaces (farms, live animal markets, abattoirs, shelters, and peri-urban wildlife corridors) to create a manageable set of sampling points that cover most exposure routes without trying to sample everywhere.

Surveillance architecture should integrate humans, animals, and environments on the same map, using subtype-aware methods. In humans, community cohorts and sentinel clinics can detect both symptomatic and asymptomatic carriage, with paired metadata on diet, travel, antibiotic use, and co-pathogens. In animals, targeted sampling of livestock, companion animals, and selected wildlife identifies which subtypes are actively moving between species. In environments, wastewater, surface waters, irrigation sources, sediment, and biofilms provide early warning and trend data. Wastewater is particularly valuable: it averages signals across large populations, provides a time series at no cost, and can be linked to community events such as heavy rain or agricultural cycles.
Study design matters as much as sampling location. Cross-sectional snapshots are suitable for mapping subtype distribution; they are bad at explaining why symptoms appear in some people and not others. Longitudinal designs, e.g., households sampled across seasons, farms sampled through production cycles, and communities sampled before and after sanitation upgrades, let you track gains, losses, persistence, and load. Pair these with dietary logs, antibiotic and antiparasitic records, and simple indicators of water and sanitation to turn noisy associations into testable hypotheses.
Diagnostics should be consistent across sectors. If humans are diagnosed by PCR, but animals and water are examined only by microscopy, comparability is lost. A minimal shared panel across human, animal, and environmental laboratories, with presence-or-absence reporting where possible, clear Ct or read-depth thresholds, and standard controls, offers benefits. When using shotgun metagenomics, publish raw reads and metadata; with targeted PCR, share primer sets, controls, and precise cycling conditions. Analytical sensitivity must be apparent because risk in water or food depends on dose and viability, not just binary detection.
Metadata acts as the glue. Every sample benefits from a concise, standard set: host species and age, clinical status, antibiotics, recent travel, diet context, co-infections, precise sampling location, date, recent rainfall, water source, and any relevant farm or processing practices. With this information, subtype-specific risk maps gain credibility. You can then explore whether STs linked to animals in a specific region also contribute to human colonisation, whether particular irrigation methods increase ST flow onto produce, or whether targeted sanitation improvements reduce community load over time.

Intervention testing fits naturally within this framework. If a watershed adds a high Blastocystis load to irrigation canals, you can assess whether upstream waste management, buffer strips, or UV treatment at key points effectively reduce environmental detection and downstream human colonisation. On farms, improved manure handling or targeted hygiene in calf pens may alter subtype profiles in both animals and workers. In clinics, standardised interpretation notes combined with watchful waiting can be compared against reflex antibiotic regimes to observe changes in colonisation, symptoms, and microbiome diversity over months rather than weeks. These are small, practical experiments embedded in real systems, not idealised trials disconnected from the exposure routes people actually experience.
Governance and ethics must be integrated from the outset. Human cohorts require clear consent for data sharing and linking; animal research needs veterinary oversight; environmental sampling should comply with local regulations and community expectations. The benefit of a One Health network is that these frameworks can be standardised and reused, thereby easing adoption at new sites and among partners. The same principle applies to equity. Field-friendly preservatives, pooled sampling, and sentinel designs enable laboratories with limited resources to participate effectively, ensuring that the global perspective is not shaped solely by well-funded urban centres.
The COST Action “Blastocystis under One Health” is poised to turn these ideas into action. It can establish the minimum metadata standards, host protocol repositories, and coordinate external quality assessments, ensuring that a positive result in a veterinary lab in rural Europe is comparable to one in a university hospital or from a river sample in a peri-urban catchment. It can also undertake the convening role that individual projects often find challenging: uniting public health agencies, water utilities, veterinary services, and academic laboratories so that surveillance, data sharing, and intervention testing are planned collaboratively rather than added on afterwards.
Finally, One Health is the ideal environment for the database and biobank needed in this field. A subtype-resolved open database that integrates human, animal, and environmental data using a unified metadata framework will enable us to track subtypes across sectors, compare interventions, and move from anecdotal evidence to identifying trends. A biobank with well-phenotyped isolates, stool aliquots, environmental samples, and blinded reference panels will enable honest and reproducible method comparisons and accelerate mechanistic research into how specific strains interact with bacteria, mucus, and bile acids. By placing these assets within a living governance framework, you create more than just publications; you develop an evidence engine that informs decisions in clinics, farms, and water management authorities.
In conclusion, One Health is not just a slogan here; it is the only framework that reflects how Blastocystis truly moves through the world. Create the map, standardise the methods, share the data, and test interventions where people and animals coexist. Doing this will stop the field from arguing over incompatible results and initiate the development of guidance practitioners have been requesting.

Why the literature conflicts – and how to fix it
Conflicting findings aren’t a failure; they’re a map for better studies. Heterogeneous diagnostics create incomparable datasets – subtype blindness blunts interpretation. Cross-sectional snapshots miss dynamics. Small cohorts inflate noise. Mechanistic work lags behind association. The fixes are clear: shared protocols and external quality assessment; subtype-resolved detection as routine; longitudinal cohorts with diet, antibiotic use, travel, and symptom diaries; larger multi-centre collaborations with harmonised phenotypes; and mechanistic studies in culture, organoids, and gnotobiotic models that link genotype to phenotype.
Where the science is headed
We are moving toward strain-resolved genomics that replaces “ST3 present” with “strain X carrying gene modules A/B/C,” allowing better predictions of colonisation and possible symptom risk. Function-first assays will quantify protease activity, glycan use, bile acid interactions, and immune modulation directly in ex vivo or model systems. Clinical reports will evolve from binary flags to risk models that combine subtype, load, host factors, and microbiome state. On the population side, environmental monitoring and targeted sanitation will be guided by subtype maps rather than total counts.
What labs and clinics can adopt now?
There are practical improvements available today. Report the method and, when feasible, the subtype; if subtyping isn’t done, say so. Use a neutral, evidence-aware interpretation paragraph with every positive result. Encourage follow-up only when symptoms persist or when other red flags are present. Contribute data – positive and negative – to shared resources so the field accumulates robust evidence rather than isolated case series. As adoption spreads, standardised language and subtype-aware reporting will lift both clinical care and research quality.

Towards a shared evidence base: database and biobank
If we want to replace debate with evidence, we need an infrastructure that outlives individual projects. Two pieces are critical.
Firstly, a public, subtype-resolved Blastocystis database should link raw reads and assemblies to comprehensive, standardised metadata: host species, age, geography, clinical phenotype, antibiotic and travel history, co-infections, diet, sequencing platform, assay targets, load estimates, and follow-up outcomes. This resource should accept both positive and negative findings, with clear data models and versioning to ensure reproducibility. It should also include simple submission tools for clinical and environmental laboratories and dashboards that enable non-specialists to explore subtype distribution and trends.
Second, a physical biobank of well-phenotyped isolates and reference materials: stool aliquots with matched metadata, cultured strains from diverse subtypes, spiked standards for assay calibration, and blinded panels for external quality assessment. This biobank would facilitate direct method comparisons, strain-level functional studies, and genuine replication across laboratories.
The COST Action “Blastocystis under One Health” serves as a natural convenor for both efforts. The network already covers expertise in human, veterinary, environmental, and methodological fields; it can define the minimal metadata set, harmonise consent and governance templates, pilot submission pipelines, and coordinate EQA rounds. A phased approach is sensible: begin with a metadata standard and a pilot repository; then add submission tools and dashboards; establish the biobank with an initial panel of reference materials; and subsequently iterate based on community feedback.
Conclusion: build the commons, together
What this really means is that progress on Blastocystis is less about choosing a side in an old debate and more about building the shared resources that allow better answers to emerge. We need people in the same room – clinicians, clinical microbiology labs, parasitologists, microbiome scientists, veterinarians, environmental microbiologists, method developers, and data stewards – working from a common playbook. The COST Action is already doing this, bringing together people across Europe and beyond; it can keep the door open, widen participation, and turn shared goals into shared infrastructure.
If we agree on standardised diagnostics and reporting, contribute to a subtype-resolved open database, and establish a biobank accessible to any serious lab, the field will cease arguing over small, incompatible studies and start building cumulative knowledge. This shift, from isolated results to a dynamic evidence base, will give clinicians more precise guidance, assist public-health teams in targeting interventions, and enable researchers to test mechanisms that genuinely matter in people and animals. The path is practical and achievable. Let’s build it together.
Acknowledgments:
This publication is based upon work from COST Action Blastocystis under One Health (CA21105), supported by COST (European Cooperation in Science and Technology).






No comments yet