Tularemia is a rare but highly infectious disease caused by Francisella tularensis, a bacterium that can evade immune defenses. Symptoms of infection can include fever, swollen lymph nodes and, in some cases, pneumonia.
What makes the pathogen especially concerning is how little it takes to cause infection — fewer than 10 bacterial cells can be enough.
Scientists at Arizona State University have taken a key step toward understanding how this bacterium survives inside the human body. For the first time, the team has isolated and studied a set of proteins that play a central role in infection, revealing a potential weakness that could eventually be targeted with new treatments.
The proteins sit within the bacterium’s inner membrane and appear to work together, forming small assemblies that help it persist inside host cells. Until now, researchers had been unable to produce and stabilize these proteins in the lab, leaving a critical part of the pathogen’s biology unexplored. By developing a method to extract and analyze them, the team has opened the door to more detailed structural studies and eventually, new ways to disrupt the infection process.
Group of proteins
The work builds on earlier studies of individual proteins, but advances the research by capturing a group of proteins that function together as a system — one that is essential for infection but had remained inaccessible.
“I am so excited about the project as we discover the structure and function of key proteins in the infection cycle,” says Petra Fromme, corresponding author of the new study. “By uncovering how these proteins come together and function, we’re starting to reveal a vulnerability in a pathogen that has been very difficult to target. This kind of insight is what lays the foundation for developing new therapeutic strategies.”
Fromme directs the Biodesign Center for Applied Structural Discovery and is joined by ASU colleagues, including first author Eranjalee Ranaweera.
The research appears in an advance online edition of the journal Biochimica et Biophysica Acta (BBA)-Biomembranes.
Potent pathogen
Tularemia is a rare but potentially serious disease caused by the bacterium Francisella tularensis. People can become infected through insect bites — often from ticks — as well as contact with animals, contaminated food or water, or even by inhaling airborne bacteria.
Once inside the body, the bacterium targets immune cells and can evade the body’s defenses. Although cases are relatively uncommon in the United States, the disease can be severe. Its ease of spread and low infectious dose have led public health agencies to classify it as a high-priority pathogen. It has also been studied in the past as a potential bioweapon.
Antibiotics can treat tularemia, but emerging resistance is a growing concern. Without prompt care, the disease can be fatal. Understanding how the bacterium infects and survives in the body is therefore critical for developing better treatments and vaccines.
Unlocking a hidden system
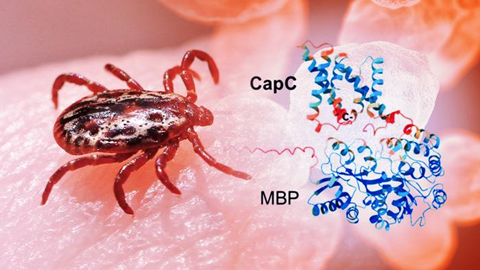
Researchers have long known that a group of proteins called the CapBCA complex is essential for the bacterium’s ability to infect and survive inside the body. These proteins are unique to F. tularensis, and animal studies show that disabling them significantly weakens the bacterium. Thus, they are a prime target for new treatment options.
But targeting these proteins has been difficult. To design drugs, scientists first need to understand what these proteins look like and how they function. Until now, that has been a major challenge because the proteins are embedded in cell membranes, making them hard to produce, isolate and study in the lab.
Cultivating the proteins
To capture and study these elusive proteins, the researchers found a way to produce them in laboratory bacteria. They inserted the relevant genes into the common bacterium E. coli. Then they added a molecular tag that helped guide the proteins into cell membranes and made them easier to detect and isolate.
Next, they used a detergent to gently extract the proteins without destroying them. The proteins were then isolated into clean, usable samples. This allowed the team to determine their shape, how they assemble into small groups and how stable they are.
Spiral proteins
The proteins were found to be mostly spiral-shaped, sticking together into small groups, rather than acting alone. The team used a combination of imaging and biochemical techniques to get the first glimpse of these proteins, revealing their overall shape and how they assemble, even though a full high-resolution structure is still to come.
“Isolating and characterizing these proteins for the first time gives us a way to visualize how the complex comes together,” Ranaweera says. “That’s a crucial first step toward understanding its function and identifying it as a potential therapeutic target, and it opens up many new directions for future work.”
The research provides the first structural clues to how this system works, laying the groundwork for deeper study and, eventually, new therapies.





No comments yet