We get to know Professor Liang Wang, who has just been appointed as new Lead Editor in Microbiology in Health and Disease at the Journal of Applied Microbiology.

The Journal of Applied Microbiology recently unveiled four key Research Themes under the JAM umbrella - Microbial Biotechnology; Systems Microbiology; Microbiology in Health & Disease; and Plant & Environmental Microbiology.
Each Theme encompasses emerging hot topics and leading-edge research that align with AMI’s goal to apply microbiology to solving the world’s greatest challenges.
Professor Wang is a full professor and distinguished medical researcher at Guangdong Provincial People’s Hospital, affiliated with the South China University of Technology and Southern Medical University in China. In a chat with The Microbiologist, he reveals what fascinates him about his new role.
What excites you about heading up this new Research Theme?
I am genuinely thrilled to lead the Microbiology in Health and Disease Research Theme for the Journal of Applied Microbiology (JAM). What excites me most is the opportunity to spotlight important research in applied microbiology, which bridges basic research with real-world challenges in how microbes influence human and animal health and disease.
In an era of emerging and re-emerging microbial pathogens, constantly increasing antimicrobial resistance, and the very complex host-microbe and host-microbiome interactions, this timely theme allows us to feature cutting-edge studies that directly contribute to improving human and animal health outcomes globally.

As Lead Editor, I really look forward to working with an international community of researchers to elevate diverse voices and foster multi-interdisciplinary collaborations. JAM’s focus on applied microbiology in health and disease perfectly aligns with my own work, and I am eager to help shape the journal’s direction by working together with JAM’s editorial team led by Prof. Andrew McBain, colleagues in Applied Microbiology International (AMI), and the fellow researchers in academic, industrial, and healthcare fields.
What first sparked your interest in microbiology and how did your career develop?
My fascination with microbiology began in childhood upon reading about the accidental discovery of the antibiotic penicillin by Scottish bacteriologist Alexander Fleming. This story sparked a deep curiosity and planted the seed of aspiration to become a scientist capable of making similar groundbreaking contributions.
During my undergraduate studies, I initially pursued theoretical physics and mechanical automation. However, later I recognized that my true passion lay in the life sciences and subsequently changed my major to biological science.

A pivotal moment occurred just before commencing my PhD, when I was offered a choice between two research projects: one focused on pure computation and the other combining computation and experimentation. I selected the latter, which allowed me to investigate how the duration of bacterial viability under abiotic stresses correlates with the evolution of bacterial virulence in both wet and dry labs. This work was conducted under the joint supervision of Prof. Michael Wise, a computer scientist, and Prof. Charlene Kahler, a microbiologist, at University of Western Australia, which has a very beautiful campus near Swan River.
Over the subsequent years, my research trajectory has evolved from foundational studies in bacterial metabolism and evolution toward more applied aspects of medical microbiology. Today, I primarily work in clinical research settings, where I apply interdisciplinary approaches to advance disease diagnosis and improve patient outcomes. This journey has seamlessly bridged academic laboratories and hospital-affiliated roles, enabling me to integrate rigorous research with tangible real-world impact.
What’s the microbe that most fascinates you and why?
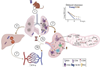
Helicobacter pylori is definitely the microbe that most fascinates me. This Gram-negative, micro-aerophilic, spiral-shaped bacterium is uniquely adapted to colonize the human gastric mucosa, thriving in the extremely acidic gastric environment through potent urease production that locally neutralizes gastric acid. H. pylori can persist in the stomach for decades with no apparent symptom for hosts. However, it is also a major causative agent of gastric diseases including chronic gastritis, peptic ulcer disease, and gastric adenocarcinoma.
The discovery of H. pylori by Professors Barry J. Marshall and J. Robin Warren, and their subsequent winning of Nobel Prize in Physiology or Medicine in 2005, fundamentally changed our understanding of infectious diseases and the treatment strategies for gastric diseases.
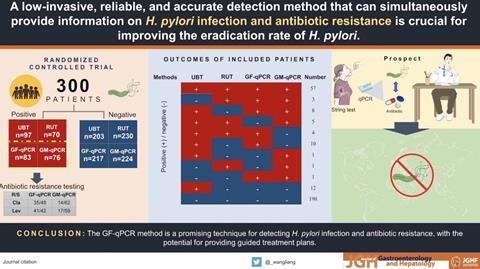
My deep fascination with H. pylori is both scientific and personal. During my doctoral studies in the Department of Microbiology and Immunology at the University of Western Australia (UWA), I trained in an academic environment that included Prof. Marshall, a full-time clinical microbiology professor there. This formative period sparked my interest and curiosity in the organism’s remarkable biology and its substantial medical significance. Later, I am honored to hold the Australia-China Helicobacter Research Fellowship, which has enabled sustained collaboration with Prof. Marshall.
A key milestone in my work was the nationwide, multi-centre, cross-sectional surveillance of H. pylori prevalence and antibiotic resistance to clarithromycin and levofloxacin in urban China, conducted in collaboration with Prof. Marshall, Dr. Alfred Tay, and the Chinese clinical partners. Using the innovative string-test technique coupled with quantitative PCR, this study provided a large-scale, contemporary data on infection rates and resistance patterns of H. pylori in urban Chinese population.

I am currently continuing this partnership with Prof. Marshall and Dr. Tay. Together, we are actively securing funding, supervising PhD students, and advancing rapid, non-invasive diagnostic platforms and precision-treatment strategies. Our shared goal is to enable accurate, resistance-guided therapy that reduces the global burden of H. pylori infection and the associated gastric diseases, with a strong focus on Asian populations where the public-health impact remains particularly heavy. The study of H. pylori will continues to drive meaningful international collaboration and innovative work that I am privileged to pursue in my academic career.
What real-world problem would you eventually love to solve with microbiology?
The real-world problem I would ultimately love to solve through microbiology is the escalating global crisis of microbial infections and antimicrobial resistance, which claims millions of lives annually and overwhelms healthcare systems. I am committed to leveraging my expertise in intelligent medicine, laboratory diagnostics, clinical microbiology, and measurement science to address this urgent challenge.

My vision centers on pioneering next-generation rapid diagnostic platforms and intelligent predictive models that enable accurate detection of microbial pathogens and rapid profiling of antibiotic resistance directly from clinical specimens. I also aspire to harness interdisciplinary approaches to identify and develop novel antimicrobial compounds, e.g., mining of traditional Chinese medicine, capable of bypassing existing resistance mechanisms at the molecular level.
Through systematic investigations into diagnostic innovation and drug discovery, this holistic strategy will advance precision prevention and effective treatment, ultimately reducing the global burden of microbial diseases and safeguarding public health for future generations.
What are the most intriguing new areas of research emerging in Microbiology in Health and Disease at the moment?
Transformative areas are reshaping microbiology in health and disease. One of the most compelling is the expanding understanding of the human microbiome and its profound influence on physiological homeostasis and pathogenesis. Microbial communities, e.g., in the gut, skin, and vagina, can modulate immunity and metabolism, with dysbiosis increasingly linked to conditions ranging from metabolic disorders and atopic dermatitis to infectious vaginitis. Advances in multi-omics and longitudinal profiling are revealing causal mechanisms, paving the way for microbiome-informed precision interventions.
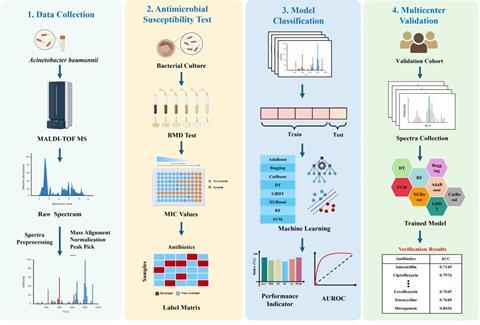
Equally promising is the rapid evolution of novel diagnostic technologies for microbial infections. Interdisciplinary integration of advanced spectroscopy, microfluidic technology, materials science, and microbial informatics is enabling ultra-fast, non-invasive, point-of-care detection and antibiotic resistance profiling directly from clinical specimens.
These platforms promise to deliver actionable results rapidly and efficiently, shifting from empirical to precision-guided therapy and significantly curbing antimicrobials overuse. These new research areas hold immense potential to mitigate the global burden of microbial diseases through earlier detection, targeted treatment, and sustainable control strategies.
How important do you think the role of applied microbiology could be in solving some of the world’s problems?
Applied microbiology plays a pivotal role in advancing SDG 3 Good Health and Well-Being, which seeks to ensure healthy lives and promote well-being for all by 2030, with key targets focused on ending epidemics of communicable microbial diseases, as well as improving access to vaccines and essential medicines.

Through applied approaches, microbiology directly enhances human health by preventing, controlling, and treating microbial-related diseases. It drives the development of improved vaccines, rapid and precise diagnostics, antimicrobial stewardship programs, and infection control strategies that effectively curb outbreaks and combat antimicrobial resistance.
These tools enable timely detection and targeted therapies, reducing the spread of resistant pathogens and supporting universal health coverage. By translating fundamental discoveries into practical interventions such as novel antimicrobials, surveillance systems, and prevention strategies, applied microbiology strengthens global health systems and delivers equitable solutions.
How would you persuade authors of the benefits of publishing in JAM?
JAM is the flagship journal of AMI and combines a rich academic heritage with high editorial standards and genuinely author-friendly policies. The journal is strongly committed to delivering an excellent publishing experience through faster turnaround times, dedicated editorial support, and a free-format submission policy that saves authors valuable time.
There are no page or colour charges for subscription-based articles, and upon acceptance, all authors receive a complimentary one-year membership to AMI, providing access to a vibrant professional community. In summary, JAM ensures that your high-quality applied microbiology research gains strong international visibility while allowing you to contribute meaningfully to the global microbiology community.
Looking in your crystal ball, what new areas of research can you see emerging in the next few years?
I foresee AI-driven novel antimicrobial drug discovery emerging as one of the most transformative areas in applied microbiology over the next few years. Generative AI will rapidly explore vast chemical and biological spaces to design entirely new classes of small molecules, antimicrobial peptides, and hybrid agents capable of bypassing existing resistance mechanisms. It will dramatically shorten discovery timelines, enabling synthesis-aware design and prediction of resistance evolution. These advances promise to deliver precision therapeutics against multidrug-resistant microbial pathogens, offering a powerful solution to the global antimicrobial resistance crisis.
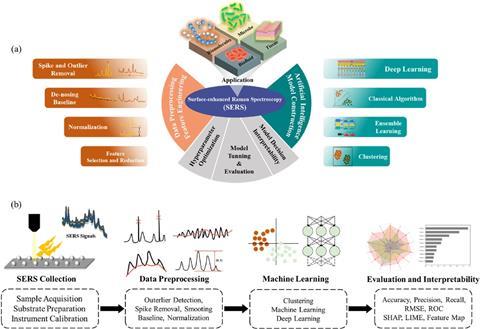
I also envision the rapid developments in microbiome research across multiple body sites as one of the most exciting frontiers in health and diseases over the next few years. Particularly promising are deeper explorations of the gut, skin and vaginal microbiomes, moving beyond descriptive profiling toward mechanistic understanding of site-specific and inter-site interactions.
Through developing personalized interventions like targeted probiotics and identifying microbiome-based biomarkers for disease diagnosis, microbiome research will significantly improve overall health outcomes.
Is there anything else you would like to add?
I would like to express my sincere appreciation to JAM and AMI team for launching the Microbiology in Health and Disease Research Theme and for the privilege of serving as Lead Editor. I encourage fellow researchers working at the interface of microbiology and human health and disease to consider submitting their best work to JAM. Together, we can advance solutions that make a tangible difference to global health. I am looking forward to engaging with the community through this theme.
Topics
- Alfred Tay
- Andrew McBain
- Antimicrobial Resistance
- Applied Microbiology International
- Artificial Intelligence & Machine Learning
- Asia & Oceania
- Bacteria
- Barry J. Marshall
- Charlene Kahler
- Clinical & Diagnostics
- Community
- Guangdong Provincial People’s Hospital
- Gut Microbiome
- Helicobacter pylori
- J. Robin Warren
- Journal of Applied Microbiology
- Liang Wang
- Michael Wise
- One Health
- penicillin
- People News
- Publishing
- Reproductive &Urinary Tract Microbiome
- Skin Microbiome/Mycobiome
- South China University of Technology
- Southern Medical University
- University of Western Australia




No comments yet