A new research paper was published in Volume 18 of Aging-US on February 8, 2026, titled “Single-cell transcriptomics reveal intrinsic and systemic T cell aging in COVID-19 and HIV.”

In this study, co-first authors Alan Tomusiak from the Buck Institute for Research on Aging and the University of Southern California, and Sierra Lore from the Buck Institute for Research on Aging and the University of Copenhagen, together with corresponding author Eric Verdin from the Buck Institute for Research on Aging, developed a new single-cell transcriptomic clock called T immune cell transcriptomic clock (Tictock) to measure aging in specific immune cells.
Immune aging increases susceptibility to infection, cancer, and chronic inflammatory disease. Most aging clocks, used to measure it, rely on bulk measurements from mixed cell populations. As a result, they cannot determine whether age-related signals reflect shifts in cell proportions or true molecular aging within defined immune cells.
To address this limitation, the research team used single-cell RNA sequencing, a method that measures gene expression in individual cells. They analyzed nearly two million immune cells from the blood of healthy adults to develop Tictock. This tool integrates automated classification of six canonical T cell subsets with cell-type specific age prediction models. This design enables the separation of systemic aging, reflected by changes in cell proportions, from intrinsic aging, which occurs within individual cells.
Impact of Covid-19
When the team applied Tictock to patients with acute COVID-19, they found two clear effects. First, COVID-19 altered T cell composition, including significant reductions in naïve CD8 and naïve CD4 T cells. Second, the infection increased the biological age of naïve CD8 T cells. In people living with HIV who were receiving long-term antiretroviral therapy, T cell proportions remained largely stable. However, naïve CD8 T cells still showed signs of accelerated aging.
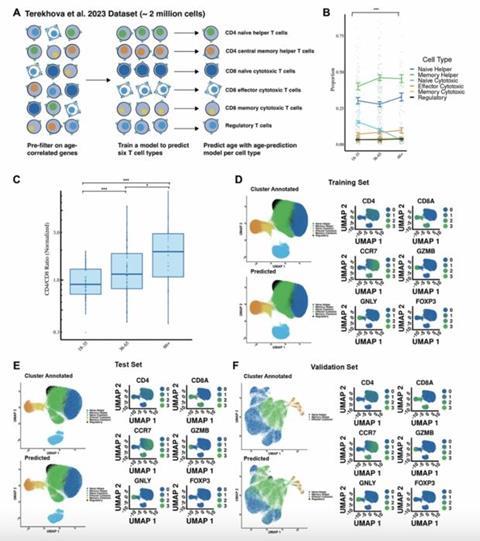
The study also uncovered shared biological pathways linked to immune aging. Many of the genes that predicted age were involved in ribosomes, the structures that help cells produce proteins. The researchers also observed that older immune cells often had shorter average transcript lengths, a feature previously linked to aging. These findings suggest that changes in protein production and gene regulation play an important role in immune decline.
Common pathways
“Gene Ontology enrichment of 209 genes shared across six clock models identified common pathways including the cytosolic small ribosomal subunit, TNF receptor binding, and cytosolic ribosome components.”
Overall, Tictock was designed to measure relative aging within defined T cell populations rather than overall biological aging. By distinguishing systemic from cell-intrinsic immune aging, it provides a clearer understanding of how viral infections such as COVID-19 and HIV reshape immune function. This approach enables the study of immune aging at single-cell resolution and may support improved immune risk assessment in clinical and research settings.
Topics
- Alan Tomusiak
- Buck Institute for Research on Aging
- Clinical & Diagnostics
- COVID-19
- Eric Verdin
- gene ontology
- immune aging
- Immunology
- Infectious Disease
- Innovation News
- Microbiological Methods
- One Health
- SARS-CoV-2
- Sierra Lore
- T immune cell transcriptomic clock
- Tictock
- University of Copenhagen
- University of Southern California
- USA & Canada





No comments yet