A new review shows that while most infections caused by the Yersinia plague bacterium can currently be treated with antibiotics, concerns about rising antimicrobial resistance mean that we need to come up with new ways to disarm the bacteria instead of killing it.
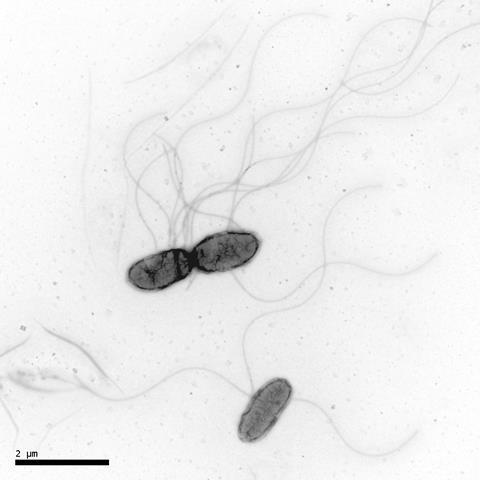
Antibiotic resistance and environmental persistence make these drugs an increasingly fragile solution and next‑generation “anti‑infective” strategies offer a more sustainable long‑term approach, warns researchers at Umeå University in Sweden.
The study, ‘Anti-infective compounds for Yersinia’ appears in Sustainable Microbiology, an Applied Microbiology International publication.
The study provides an up‑to‑date synthesis of global research into how Yersinia bacteria cause disease and how we might control these infections without relying solely on traditional antibiotics. The review was written in response to growing concerns about antimicrobial resistance, climate‑driven disease emergence, and food‑chain transmission of Yersinia species affecting both humans and animals, said corresponding author Professor Matthew Francis.
Resistant strains
“Yersinia bacteria cause a wide range of diseases, from common food‑borne gastrointestinal infections to plague, and they circulate between humans, animals, food systems, and the environment. While many infections are treatable today, resistant strains are emerging, and these bacteria are particularly adept at persisting in soil, water, food systems, and animal reservoirs, where resistance genes can spread silently,” he said.
“The core problem we address is that future outbreaks may become harder to control if we continue to rely on antibiotics alone. We wanted to understand whether alternative strategies – those that block virulence, transmission, or host damage rather than killing the bacteria outright – could provide safer and more sustainable solutions.”
The review systematically examined decades of peer‑reviewed research to understand how different Yersinia species cause disease and which of their biological processes might be realistically targeted to control infection.
Rather than focusing on individual compounds, the team concentrated on identifying core virulence mechanisms, genetic regulatory circuits, and lifestyle switches that allow these bacteria to survive, spread, and evade host defences.
Switching between lifestyles
They describe how Yersinia tightly controls its behaviour through gene regulation, enabling it to switch between distinct lifestyles, for example, environmental survival, transmission, and active infection of a host. By mapping these regulatory constraints, they identify points of vulnerability where intervention could disarm the pathogen without necessarily killing it. Key examples include secretion systems used to inject toxins into host cells, regulatory pathways that coordinate stress responses and biofilm formation, and signalling systems that control adhesion and immune evasion.
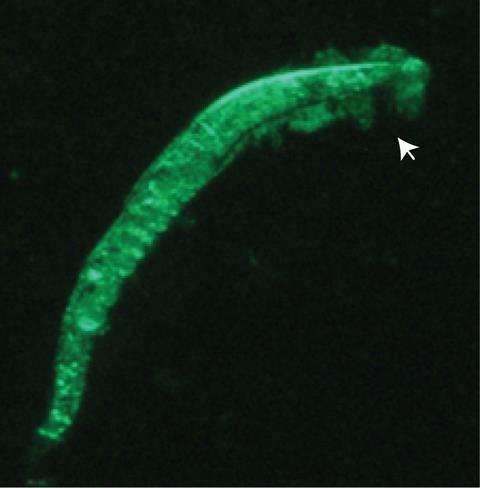
The review then outlines how these targets have been experimentally inhibited in basic research using approaches such as small‑molecule inhibitors, antibodies, bacteriophages, vaccines, probiotics, and host‑directed strategies. For each approach, it summarises what happens when a specific system is blocked, for example, reduced immune suppression, impaired transmission, or loss of virulence in infection models, and critically assess the limitations of these studies, including off‑target effects, lack of efficacy in vivo, evolutionary compensation by the bacteria, and obstacles to clinical translation.
No single strategy
Together, this analysis shows that while no single strategy is sufficient on its own, understanding virulence control, regulation, and lifestyle choice provides a rational framework for designing more sustainable anti‑infective therapies.
“One of the most striking findings is how much innovation exists in the laboratory, yet how little of it has reached real‑world use. Despite strong experimental results, no dedicated anti‑virulence drug for Yersinia has yet made it into clinical practice,” noted Professor Francis.
“We were also struck by how adaptable Yersinia bacteria are. When a major virulence mechanism is blocked, the bacteria can sometimes compensate by activating alternative pathways, meaning that inhibiting a single system may not always be sufficient to disarm the pathogen.
Reprogramming under stress
“What is especially surprising is how little we still know about how Yersinia selects, regulates, and prioritises these compensatory responses. Despite decades of detailed knowledge about Yersinia virulence and gene regulation, the rules governing how these bacteria reprogramme themselves under stress remain poorly understood. This is particularly true in the context of exposure to anti‑virulence inhibitors, where selective pressure does not kill the bacteria but instead encourages physiological adaptation.”
These gaps highlight that much of Yersinia biology remains unexplored, especially when it comes to understanding compensatory evolution in response to therapeutic interventions, which is knowledge that will be essential if anti‑infective strategies are to succeed in the long term.
“This work reframes how we think about treating bacterial infections. Rather than escalating the antibiotic arms race, anti‑infective strategies aim to disarm pathogens instead of killing them, reducing selective pressure for resistance and helping to preserve existing antibiotics for situations where they are genuinely indispensable,” Professor Francis said.
Beyond clinical medicine
The implications extend well beyond clinical medicine. Yersinia species circulate between humans, animals, food systems, and the environment, making them highly relevant to food safety, aquaculture, climate resilience, and global health security. Reducing antibiotic use in agriculture, strengthening non‑antibiotic disease control in aquaculture, and limiting environmental reservoirs of Yersinia can all help slow the spread of antimicrobial resistance while protecting ecosystems and food supplies.
“Importantly, what we learn from Yersinia, including both the strengths and limitations of anti‑virulence and alternative control strategies, can inform efforts against other bacterial pathogens where antibiotic resistance is already a severe clinical problem. In this sense, Yersinia serves not only as a pathogen of concern but also as a valuable model for developing more sustainable approaches to infectious‑disease control more broadly,” Professor Francis said.
The way forward
Future progress depends on integrated strategies. Promising compounds must be tested earlier in realistic infection models, combined with antibiotics or immune‑based therapies, and designed with drug delivery and safety in mind from the outset.
Equally important is policy and funding support for diseases that are dangerous but relatively rare, such as plague, where commercial incentives are limited. Without coordinated international investment, many sound ideas will remain stuck in the laboratory.
The review was led by Kumar D. Gahlot, Matthew S. Francis, and Jyoti M. Gurung at the Department of Molecular Biology and the Umeå Centre for Microbial Research, Umeå University, Sweden.
Research in the Francis laboratory has been supported in part by the Swedish Research Council, the Foundation for Medical Research at Umeå University, and the Faculties of Science and Technology and Medicine at Umeå University.
The study is published in Sustainable Microbiology.
Topics
- AMR in the Environment
- Antibiotics
- Antimicrobial Resistance
- Applied Microbiology International
- Bacteria
- Biofilms
- Community
- Disease X
- Emerging Threats & Epidemiology
- Food Security
- gene regulation
- Infection Prevention & Control
- Infectious Disease
- Jyoti M. Gurung
- Kumar D. Gahlot
- Matthew Francis
- One Health
- Pharmaceutical Microbiology
- Research News
- UK & Rest of Europe
- Umeå University
- Yersinia






No comments yet