Since a landmark 2009 study, researchers have known that a common gut bacterium, Bacteroides fragilis, drives colon tumor formation, potentially leading to colorectal cancer, by secreting a toxin that damages the lining of the colon. But until now, the exact mechanism the toxin uses to latch onto those cells remained a mystery.
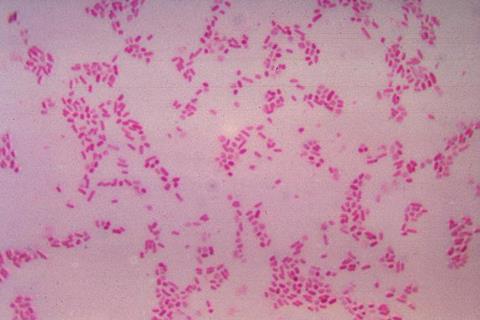
A multi-institutional team led by researchers at the Johns Hopkins Kimmel Cancer Center Bloomberg~Kimmel Institute for Cancer Immunotherapy and the Johns Hopkins University School of Medicine has identified the missing link. The study, published April 22 in Nature, reveals that the B. fragilis toxin BFT must first bind host receptor claudin-4 before it can cause damage. The work was supported in part by the National Institutes of Health.
“We’ve made several attempts over time to identify the receptor, so this is an exciting moment,” says senior author Cynthia Sears, M.D., Bloomberg~Kimmel Professor of Cancer Immunotherapy and professor of medicine at Johns Hopkins. “Understanding how bacterial toxins work can open doors to new approaches for detection and therapy for associated diseases, including diarrhea, colorectal cancer and bloodstream infections.”
The discovery, in fact, already led to the development of a molecular decoy that successfully blocked the toxin’s effects in animal models, offering a potential strategy for preventing B. fragilis damage to the colon.
Potent ability
B. fragilis can be detected in up to 20% of healthy individuals, and has a potent ability to induce colon inflammation and tumor formation. Prior work in Sears’ lab showed that BFT triggers chronic inflammation in the gut by dividing E-cadherin, a protein essential for maintaining the colon’s protective barrier. In their earlier Nature Medicine study, Sears and colleagues showed that the action of BFT leads to colon tumor formation. Yet BFT did not appear to directly attach to E-cadherin. Some other, elusive mechanism appeared to be at play.
Identifying that mechanism began with a genomewide CRISPR screen led by Maxwell White, an M.D./Ph.D. candidate in the Sears lab, in collaboration with the lab of Matthew Waldor at Harvard Medical School. By systematically knocking out genes in colon epithelial cells, White and colleagues from Waldor’s lab identified claudin-4 as the link. When White knocked out claudin-4, BFT toxin was unable to bind to the cells, leaving the E-cadherin target untouched.
“It took a while to get the assay working and validate the approach, but once we were able to do the screen, claudin-4 was a clear, resounding top hit,” says White. “That was an exciting moment.”
Surprising reveal
The identity of the receptor was a surprise, adds Sears, as she and others in the field had long expected the receptor to be a signaling protein, such as a G-coupled protein receptor, which claudin-4 is not. In a literature review, the team could not identify any other toxin that functions this way, as most proteases go straight to their targets rather than binding a separate receptor first.
To confirm that the toxin and the receptor were physically locking together, the Johns Hopkins team partnered with structural biologists F. Xavier Gomis-Rüth and Ulrich Eckhard at the Molecular Biology Institute of Barcelona. Using biophysical analysis, White and the Barcelona group demonstrated that BFT and claudin-4 form a tight, one-to-one complex in a test tube, providing the first physical evidence of the binding interaction.
The research then moved from the test tube into living systems through a collaboration with the lab of Min Dong at Harvard Medical School. With Kang Wang and colleagues in Dong’s lab, the team evaluated how the toxin behaved in the complex environment of the gut using mouse models.
Decoy receptor
By creating a decoy claudin-4 — a soluble protein that displayed claudin-4 sequences — the researchers attempted to prevent the toxin from binding colon cells. Indeed, BFT bound to the decoys instead of the receptor. This decoy strategy successfully protected mice from BFT-induced damage.
“This approach could be iterated upon with small molecules or other biologics that have better pharmacological properties,” says White. The team is now exploring which molecular approaches might be most successful to block the toxin.
The researchers note that one piece of the puzzle remains: While the team identified the receptor and proved the binding, the exact experimental structure of the interaction between BFT and claudin-4 has yet to be captured. Current AI modeling tools, such as AlphaFold, were not able to fully resolve the interaction.
Additional authors on the paper include Jason Chen, Shaoguang Wu, Abby L. Geis and Jessica Queen at Johns Hopkins and Hailong Zhang, Karthik Hullahalli and Jie Zhang at Harvard Medical School.
The research was supported by the Bloomberg~Kimmel Institute for Cancer Immunotherapy, Janssen Research and Development, Cancer Research UK, the National Institutes of Health (grant numbers R01 AI042347, R01 NS080833, R01 NS117626, R01 AI170835 and R01 AI189789) and the Howard Hughes Medical Institute.
Sears receives royalties for writing and reviewing for UpToDate. This arrangement is managed by The Johns Hopkins University in accordance with its conflict-of-interest policies.
Topics
- Bacteria
- Bacteroides fragilis
- Cancer Microbiology
- claudin-4
- colon tumor formation
- Cynthia Sears
- Disease Treatment & Prevention
- E-cadherin
- F. Xavier Gomis-Rüth
- Gut Microbiome
- Harvard Medical School
- Immunology
- Infection Prevention & Control
- Johns Hopkins University
- Kang Wang
- Matthew Waldor
- Maxwell White
- Min Dong
- Molecular Biology Institute of Barcelona
- One Health
- Research News
- Structural Biology
- Ulrich Eckhard
- USA & Canada





No comments yet